Cornell structural biologists adopted a new strategy for using a great technique for X-ray analysis to catch something the conventional strategy had never represented: the collective movement of proteins. Also, they did as such by making software to painstakingly stitch together the scraps of information that are usually dismissed simultaneously.
Their paper, “Diffuse X-ray Scattering from Correlated Motions in a Protein Crystal,” published March 9 in Nature Communications.
As a structural biologist, Nozomi Ando, M.S. ’04, Ph.D. ’08, assistant professor of chemistry and chemical biology, is keen on charting the movement of proteins, and their internal parts, to more readily comprehend protein function. This sort of movement is notable yet has been hard to document because the standard procedure for imaging proteins is X-ray crystallography, which produces essentially static snapshots.
“Because we’re studying challenging biological systems, the group often has to pioneer new structural methods as well,” said postdoctoral researcher Steve Meisburger, Ph.D. ’14, the paper’s lead author. “One of the questions that we have been interested in since the beginning is how a protein’s subtle breathing motions direct biochemical function.”
The analysts carried their project to the Cornell High Energy Synchrotron Source (CHESS), where they took advantage of the facility’s Pilatus 6M pixel-array detector, which empowered them to make extremely high-resolution pictures.
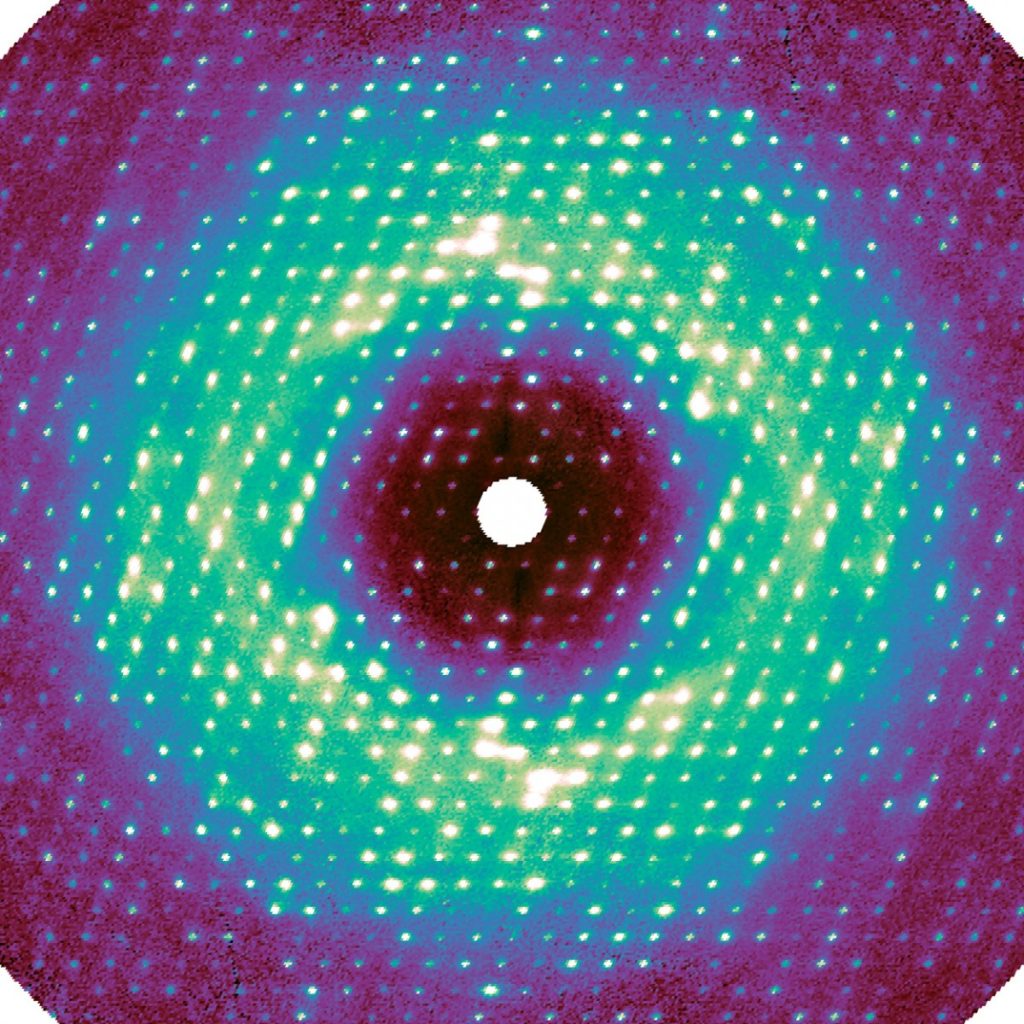
For this work, as in regular crystallography, X-rays were radiated at a sample crystal. The pixel-array detector recorded the intensity of the X-rays that were diffracted by the crystal’s proteins, in this way encoding the atomic structure. Any disorder—i.e., movement—inside the crystal made extra photons bounce out, making a very weak background signal called diffuse scattering. This data has been generally discarded during data processing.
“The photons go everywhere, and the signal appears extremely weak because it’s spread out,” said Ando, the paper’s senior author. “For decades, people couldn’t measure it accurately, and they didn’t know how to interpret it.”
Meisburger made software to process the roughly 50 million one of a kind information points, bringing about a high-quality three-dimensional map. A lot to the analysts’ astonishment, the map uncovered that a critical component of this diffuse scattering pattern was an aftereffect of the protein lattice vibrating. This jiggling movement was so dominant, it appeared to obscure any movement inside the proteins, which was at first failure for the scientists.
In any case, in the wake of representing these lattice vibrations in simulations, the scientists distinguished internal protein movements too. These movements incorporated the opening and closing of the active site of the protein.
“Imagine the crystal being like a row of people trying to walk together while holding hands, but at the same time, each individual might be doing something slightly different,” Ando said. “The signal from everybody moving together is dominant, so we couldn’t discern the subtle signal that was coming from the individuals. That was something that had never been accounted for.”
This new way to deal with diffuse scattering could assist analysts with getting a more clear image of protein structure and dynamics and, at last, a superior comprehension of biochemical responses.
“We want to push this in a direction where lots of people can use the technique and learn something new about their protein,” Meisburger said. “One great thing about it is you get diffuse scattering for free anytime you do a regular crystallography experiment. This technique adds information to what you would normally get.”
Lee Triebel is a famous author and has written even more famous books based on his own experiences. At the age of 20, he started his career in writing. His Bachelor’s degree in English literature helped him render his passion. He is one of the reputed writers. Now he is working on the Herald Quest website as a freelance author.
Disclaimer: The views, suggestions, and opinions expressed here are the sole responsibility of the experts. No Herald Quest journalist was involved in the writing and production of this article.